scBench Updates: Opus 4.7, GPT 5.5, Gemini 3.1
Benchmarking frontier models on messy, real-world single-cell data analysis
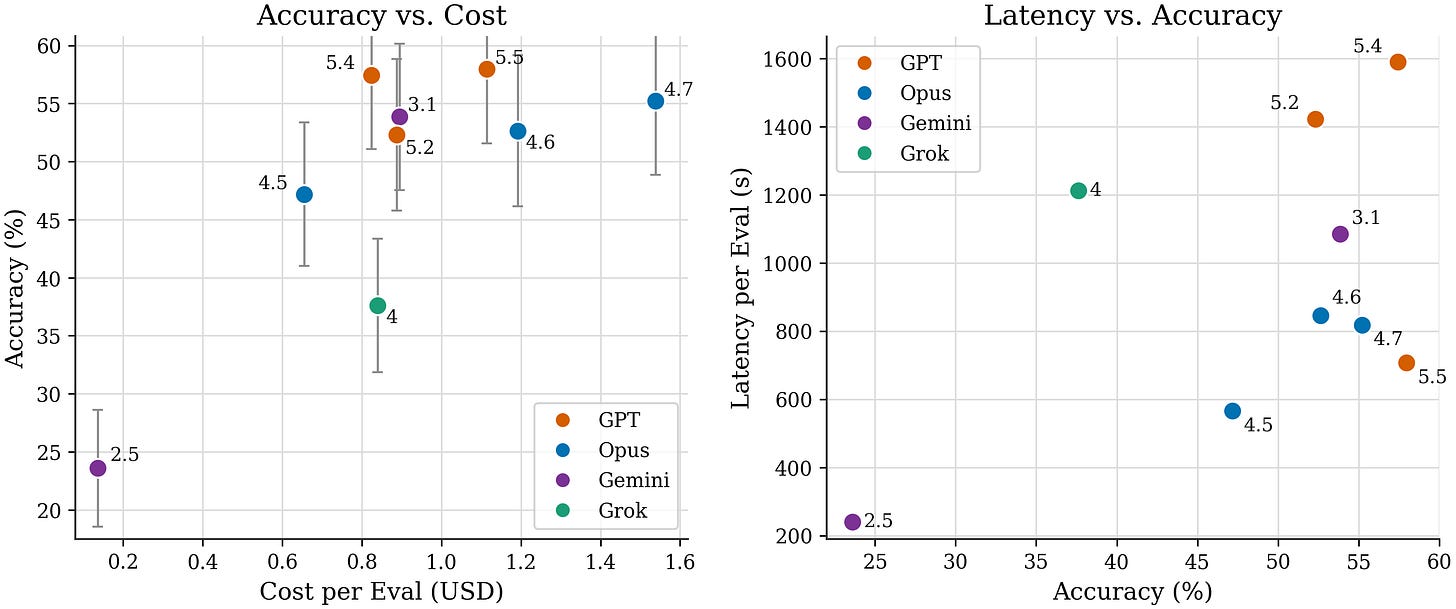
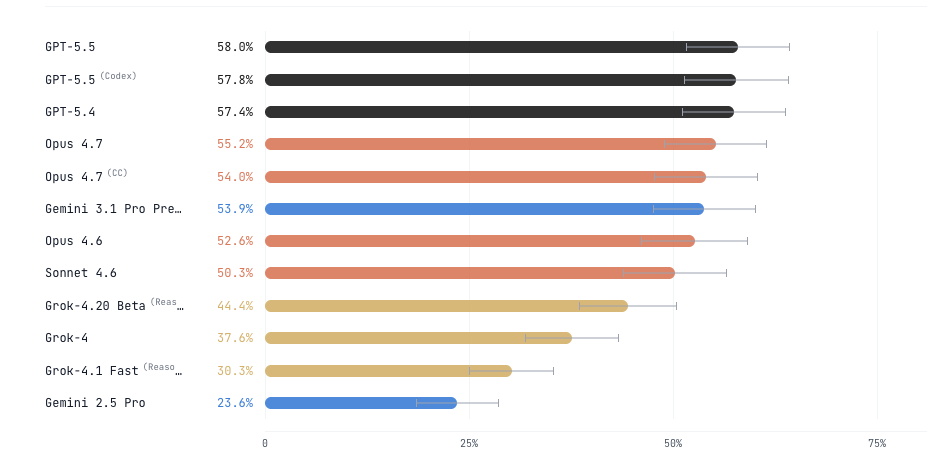
We revised scBench with additional rounds of scientist review and generated data for new models like Opus 4.7, GPT 5.5, and Gemini 3.1.
On accuracy, Gemini 3.1 doubles, Opus 4.7 shows modest improvement and GPT 5.5 shows no significant improvement.
On latency, GPT 5.5 improves greatly, Opus 4.7 shows no significant improvement and Gemini 3.1 worsens by ~5x.
Full data and example evals/trajectories available on benchmarks.bio.
Models are showing steady improvement in generic data analysis ability
Trajectory review reveals improvement in exploratory behavior:
Thoroughly investigating data and thinking through experimental context before picking which analysis algorithm or tool to use
Self-verification after finishing steps or submitting answers
Early signs of assay-aware analysis steps (eg. QC thresholds) specific to kit/machine type instead of generic tutorial steps. Largely explains the jump in Opus 4.7 accuracy.
Models still struggle with biology-specific analysis details
Many remaining evals require knowledge of experimental design, kit/machine-specific analysis details, or tacit scientific knowledge. Some examples:
Aggregating to the correct statistical unit before testing (ex: inappropriately treating cells as statistical replicates, inflating power in statistical tests)
Applying whole transcriptome scRNA-seq defaults to targeted 30-400 gene panels (ex: BD Rhapsody)
Accurately recognizing and removing depth/batch confounds (ex: PC1 absorbs library size)
Integrating data from multiple platforms (ex: DNA + RNA on Tapestri, protein + mRNA)
Explore data and trajectories on benchmarks.bio
We regularly update our benchmark family with new models: benchmarks.bio. Encourage those interested in understanding what these benchmarks actually measure to inspect sample tasks and trajectories.