Agentic biology is shaped like software
Biology will not jump straight to autonomous AI scientists. Like software, it will first accelerate where work is executable, feedback-rich, and economically bottlenecked: data analysis.
Introduction
It’s important to remember just how surprising the rise of software engineering agents was. A year or two ago, most professional engineers thought of AI tools as code generation assistants - useful for formatting, refactoring, or scaffolding unit tests - but never capable of grokking complex requirements, modifying large codebases, or making systems-level engineering decisions.
Today, 4% of all public GitHub commits are authored by Claude Code and Mythos is uncovering thousands of day-zero vulnerabilities *autonomously* in workhorse software. Agents build browsers and compilers. Many of my most skeptical peers now use multiple agents in parallel to tackle daily, high stakes work.
In hindsight, many people treat this capability growth as inevitable. Software, after all, is “obviously verifiable”: unlike other domains grounded in the messy, physical world, code runs deterministically on computers. But software is more than code and frontier agents are starting to show higher-level engineering judgment on open ended tasks.
I believe biology will follow a similar path. Research workflows in biology will reorganize around agents for the same reason software did: both domains contain a layer of work that is concrete, executable and feedback-rich. In software, that layer is code. In biology, its measurement-grounded data analysis.
From this analogy, we’ll build up three claims using real examples:
First, the first useful biology agents will be data-analysis agents, not autonomous scientists, just as coding assistants came before increasingly autonomous engineering agents.
Second, assay-specific data analysis is needed for true biological reasoning to emerge.
Third, the analysis layer becomes economically more important as molecular data generation grows.
Software fell first because code is executable and feedback-rich
To understand what I mean by “both domains have a layer of work that looks the same”, let’s first dig into what that means for software.
Software is written in code, which has some nice properties. You can run it quickly through a compiler and get outputs and errors. You can easily inspect the intermediate state of the digital objects you build up in a program by printing values or writing them to files.
Code is then an ideal substrate for agents to learn tasks. They can write and run chunks, inspect outputs and loop. But the useful things we build in software are far more than just code. They incorporate subjective and somewhat open-ended engineering decisions, many inherited from the domain in which they’re used and the problems they solve.
It is then surprising agents developed the capability to autonomously construct, for example, browsers. “The verifiability of code” does not explain the progression from passing tests to making nontrivial systems decisions about concurrency, caching, latency and security.
The obvious reason is skill in higher-order concepts emerged after focused training on the procedural skill of code is because you have to get to the parts of the problem where this knowledge is useful. It’s difficult to learn how to rate limit an endpoint if you struggle with Python function syntax or are unable to reason correctly about event loops.
But more subtly, and important to the analogy with biology, is detailed knowledge of your building material informs the higher-level decisions. It’s difficult to make good browser-level decisions (caching, rendering) without understanding the lower-level behavior of code.
Biology has a similar analytical substrate
So what does any of this have to do with biology?
Most modern biology papers are centered around large-scale measurement experiments and share the following structure:
Choose a biological model (cell line, animal, organoid) and variables/controls
Generate data from the model
Process the data
Creatively thinking about the analytic results in context of prior literature
Make scientific claim
If you trace how an agent might work through this flow to tackle research autonomously, I argue:
Analysis of assay-specific data analysis is necessary to reach stages where scientific reasoning is useful
Scientific reasoning tasks are shaped like high level software engineering tasks, both in complexity and their dependence on underlying procedural skill in data analysis for accuracy.
So the existence of a thick underlying layer of molecular data analysis scaffolds scientific thinking in a similar way that code scaffolds more complex software engineering judgement.
The best way to make this concrete is with an example.
An example: reprogramming fibroblasts into brain organoids
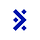
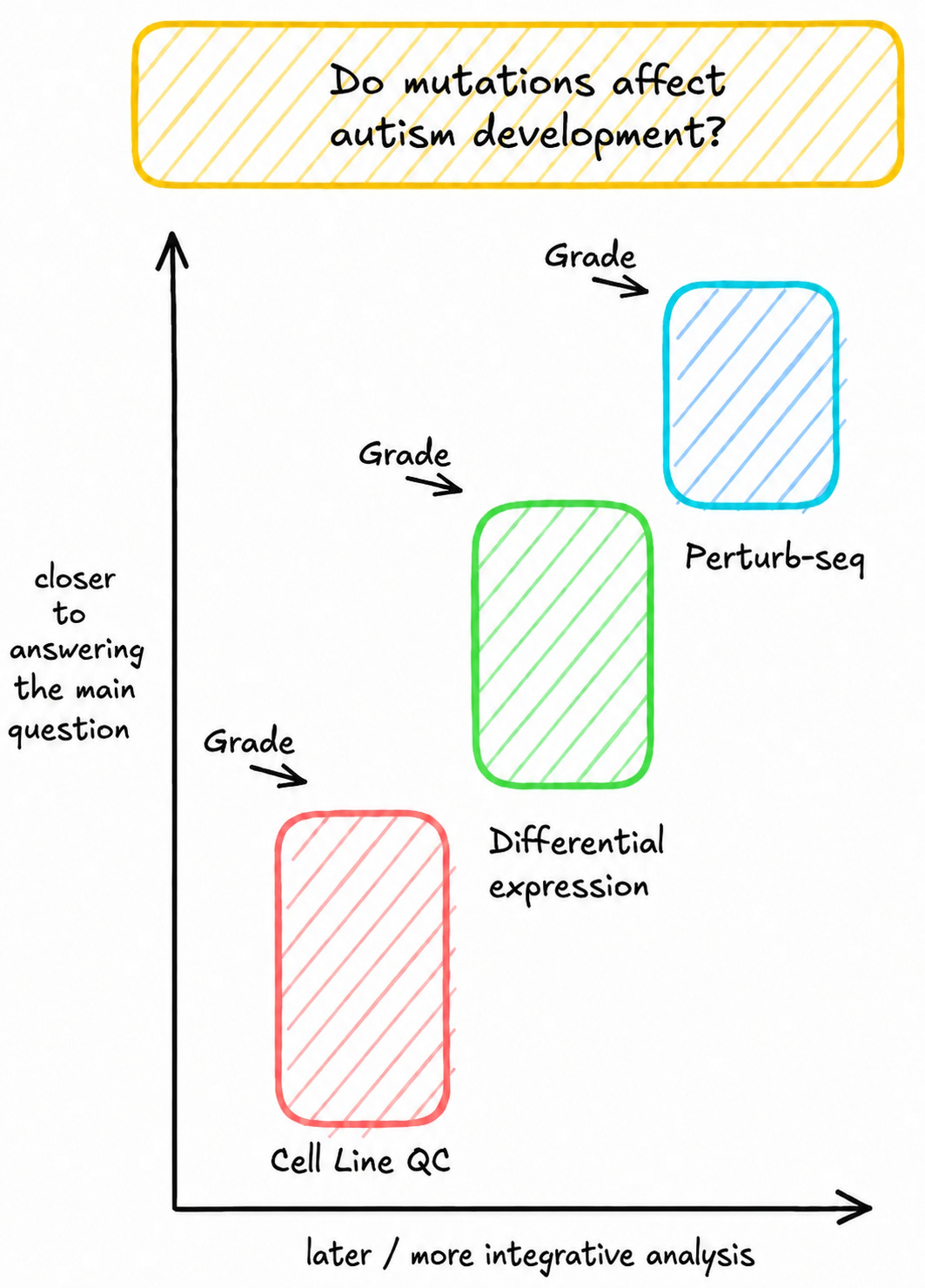
Let’s take a look at a recent study from Gordon et al. where authors harvest fibroblasts (very common ‘connective’ cells) from 55 autism patients, reprogram them into stem cells with Yamanaka factors and turn each stem cell into a brain organoid. Their central question is “do the different mutations in each patient affect the development of autism”.
This is a good example of the type of project (structure, scale, difficulty) we might want an agent to tackle in the future.
We can immediately see the question admits no “global verifiability”. It’s unclear how to produce a correct value to grade against (or what type of value would even make sense). A ground truth would depend on dozens of intermediate observations, some without clear scientific consensus: how organoids are quality-controlled, how cell types are annotated, how gene expression modules are defined.
But we can decompose the paper into smaller steps with clearer edges, some even with “locally verifiable” results.
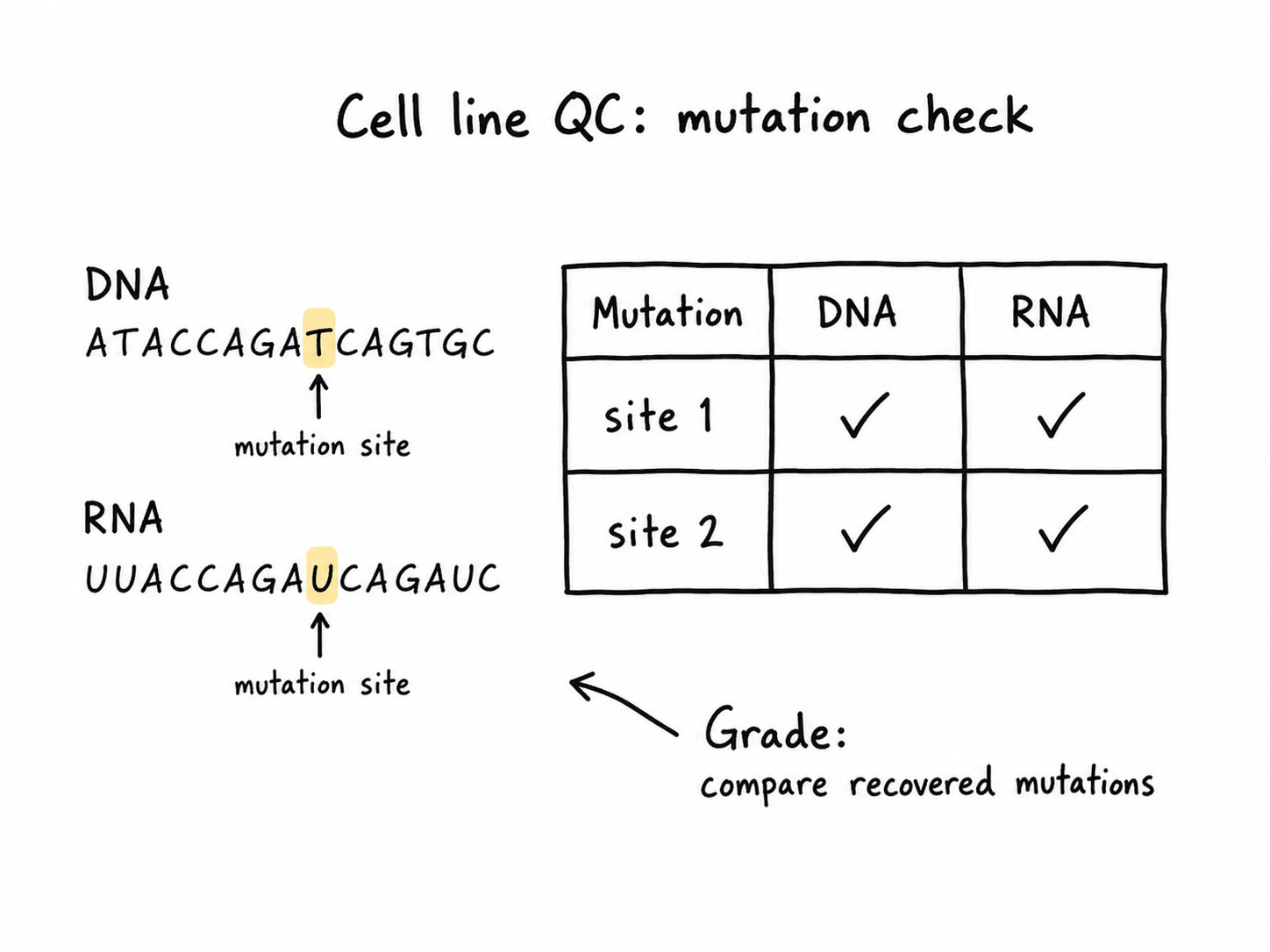
Cell line QC
First, the agent compares mutations in whole-genome sequencing and RNA-seq data to quality control stem cells.
We grade the agent on which cell lines it drops and which mutations it recovers for each line.
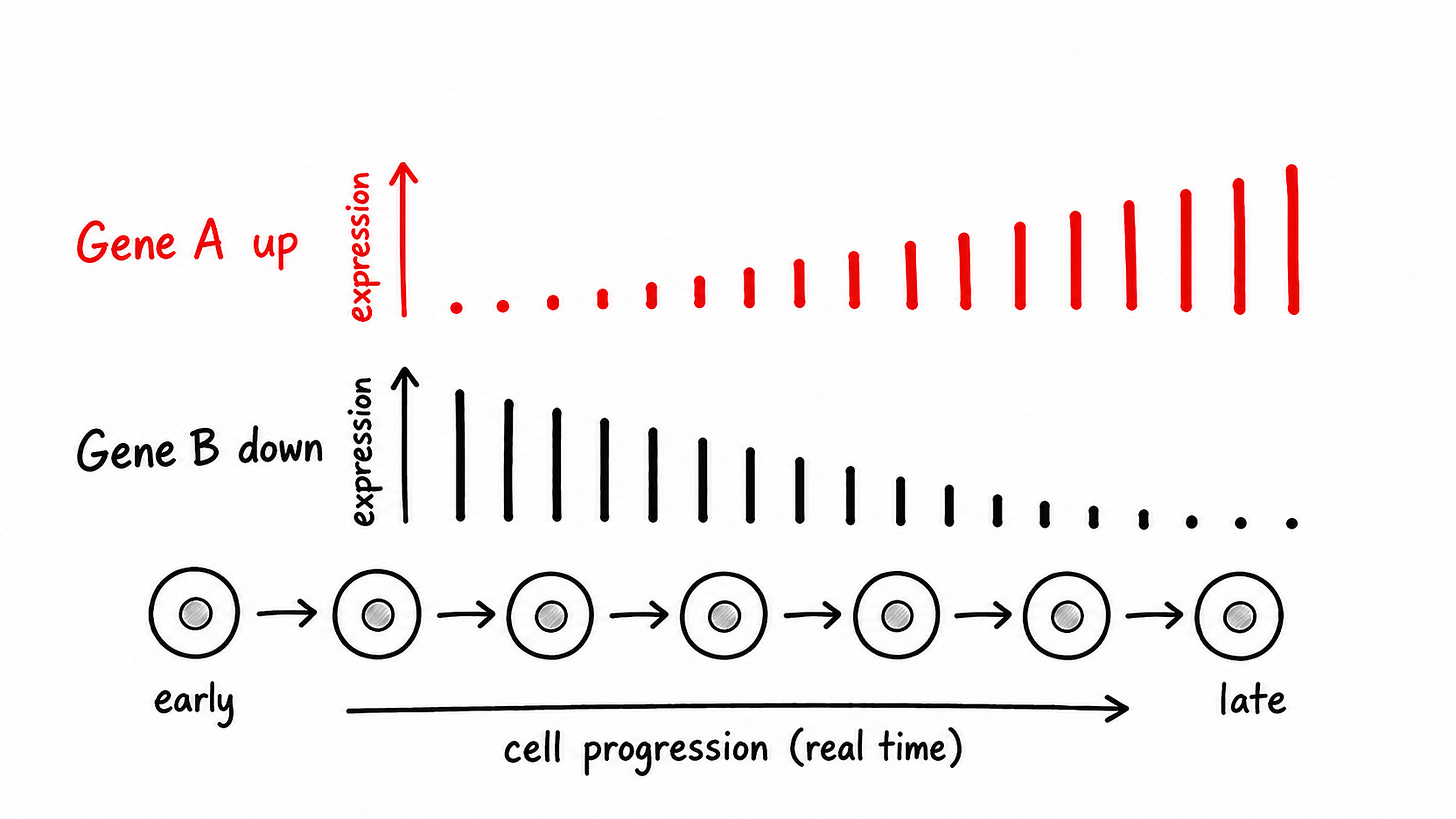
Differential expression
Then, the agent looks across cell lines to recover genes that shift the most over time (as the cells develop).
We grade the agent on the agreement with the genes recovered by the author.
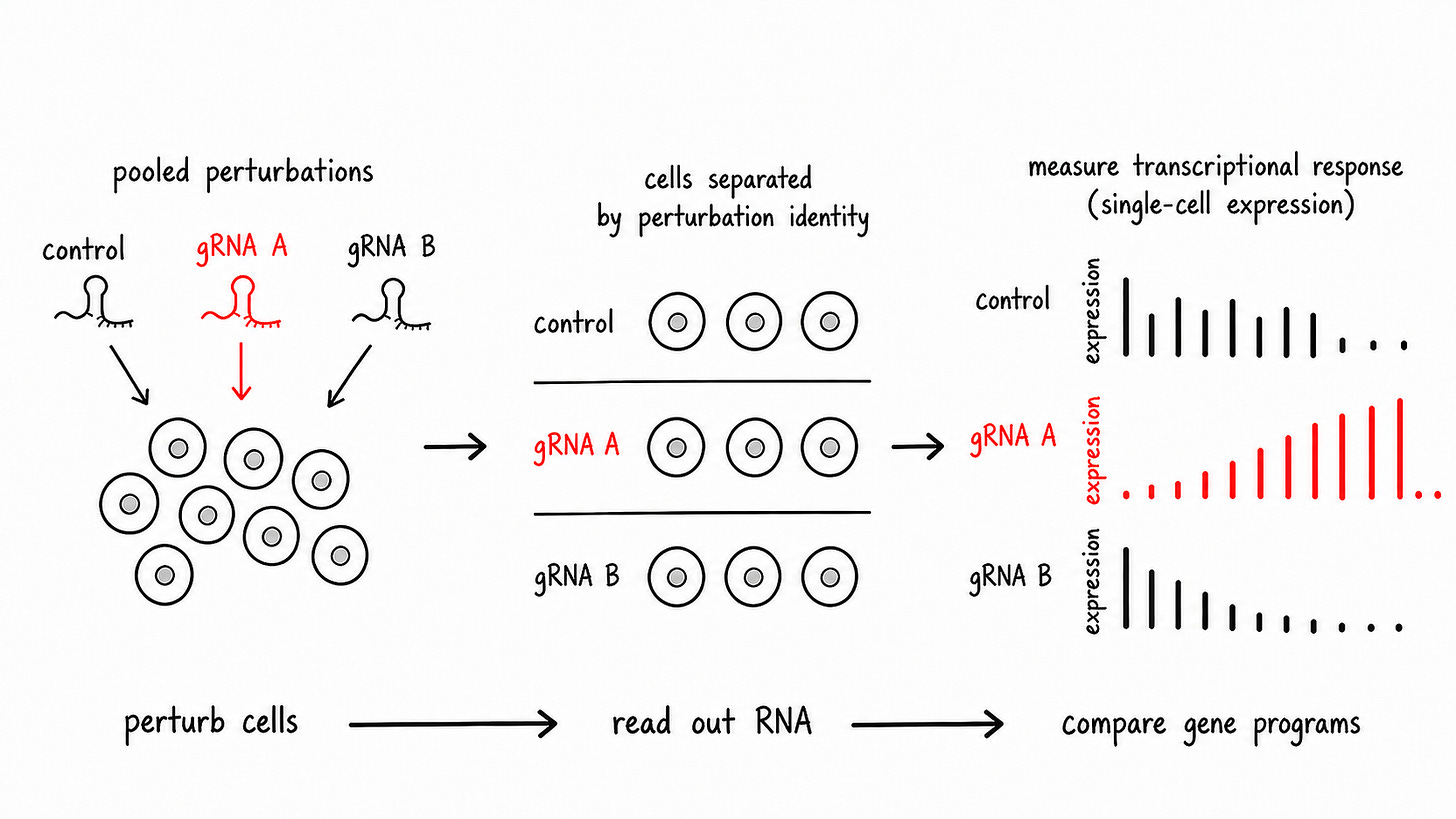
Perturb-seq
We then use CRISPRi to actually “knock down” these genes in the lab. The agent analyzes this Perturb-seq data to identify which of these genes had a functional effect when it was repressed.
We grade the agent on the genes the authors concluded were important.
Each of these analysis steps start from raw measurement data and build up to a well-defined scientific outcome. All of them are necessary for a scientist to draw a conclusion about the overarching question.
Data analysis is the gateway to scientific reasoning
As shown above, agents can decompose biological research into concrete analysis steps, with each step building upon the previous one. Let’s now get a sense of the higher-level tasks remaining after the agent works through the data:
Compare data from different assay types to validate results: bulk + sc-RNA-seq, IP-MS + Perturb-seq
Compare expression across technical (sequencing lane, differentiation batch) and biological (donor, cell line) covariates
Scour existing literature for prior biological evidence (eg. known disease associations or functions of candidate genes)
These tasks are much more “AI Scientist” shaped. They require synthesis across modalities, judgment under uncertainty, and knowledge of prior biology. But they are not obviously more complex than the kinds of open-ended engineering judgment agents are beginning to perform in software. The practical bar is matching the judgment of a strong scientist: reasoning from the available measurements, controls, analysis choices, and literature to the conclusion best supported by the evidence.
And because this biological judgement is empirical, the agent must reason from noisy data and understand analysis assumptions well. Developing scientific judgement then requires mastery of the analysis steps and cannot be separated from it.
For instance, an agent would need to understand the statistical choices made in the differential expression analysis. Decisions like which samples are included, how genes are filtered, how covariates (like sequencing lanes) are factored out can modify the results. An agent could conclude “a gene is downregulated” because these genes genuinely co-vary across mutation or because the gene is capturing a hidden variable in the data: developmental maturity, cell-type composition, sequencing depth, some temperature in the lab, etc.
Another example is how metadata is constructed. If the agent is looking for genes that regulate development, the exact way it constructed the cell type labels in the data need to be understood. Depending on how we define “cells in early development”, low count genes in this population could surface from genuine biology or an incorrect definition of this population with subjective marker genes.
An agent with an understanding of these assumptions is equipped to make accurate scientific decisions. If it did the analysis itself, it also has these details fresh in context.
Where the analogy breaks
Biology is among the last frontiers of the empirical sciences. The ‘objects’ of study (living organisms) span angstrom (protein) to micron (cell) to meter (leg) scale, synthesizing phenomena across chemistry and physics. The measurements we use to study them are produced by instruments and protocols on the frontier of human knowledge with noise and error.
This complexity means the analogy to software breaks in at least two places.
First, biological ground truth is more difficult to define. Even apparently well-scoped tasks like clustering cells, annotating cell types or calling differentially expressed genes rarely have one canonical answer. This is an active area of research, with ongoing work to develop practical methods and tools to constrain ambiguity and model scientific intent verifiably. It remains very difficult.
Second, feedback is much weaker once we leave the analysis layer. For data analysis, the feedback loop still looks somewhat like software: run code, inspect outputs, loop. However, true feedback for higher order scientific reasoning requires some control over the data-generating process. We will likely make a lot of progress using ground truths constructed and graded a lot like code without control over this loop. But true autonomous science requires direct feedback with the substrate being studied.
This layer becomes more important as biology scales
Agents will need to develop data analysis capability first (at least for real work) and accurate scientific reasoning seems in part dependent on a detailed understanding of analysis. It’s important to now understand how much more important analysis might be in the context of data generation trends.
Molecular data generation follows an exponential curve and analysis costs are rapidly becoming more expensive than reagents/prep labor with new assay generations. An increasing proportion of biological work now happens after the experiment has finished in the lab.
If you play out these trends, the bottleneck will shift from producing measurements to interpreting them. The question is not if but when. Future datasets will dwarf the stock of past biological data, and human capacity to analyze them will not scale at the same rate. The answer to this will be agents spinning in harnesses that reliably take data to scientific conclusions, deployed in R&D workflows across biotech where data is bottlenecked today.
The surface area of computational biology is growing
People often equate ‘computational biology’ with narrow analysis tools, like sequence assembly or differential expression software. While these have been important contributions of the field, computational biology should really be thought of as “the broad use of computers to solve problems in biology”.
For years, the components have been building for a computational shift in biology: rapidly compounding molecular data, increasingly automated labs, and iterative design-build-test-learn workflows. Yet these advances have not translated into an obvious step-change in aggregate R&D productivity.
That now appears to be changing. The industry is reorganizing around AI and computing, in how experiments are designed, data is analyzed, companies are formed, and pharma structures partnerships. Computers are becoming the interface through which agents “see”, understand and eventually manipulate living systems.
The first useful deployment of these agents will not be autonomous AI scientists but analysis collaborators embedded in real workflows. This is both where true need lies and what they will get good at first. Biology has no shortage of interesting new ideas. The bulk of economically valuable agentic work is concentrated where computers can make progress on R&D cycles, augmenting scientists where they are today.
Thank you to Aidan Abdulali, Anirudh Narsipur, Brandon White, Kyle Giffin, Alfredo Andere, Aashay Sanghvi, David Yang, Hannah Le, Lada Nuzhna, Tim Proctor for revising and/or contributing to the thinking in this essay